Abstract
Recurrent urinary tract infections (rUTIs) are a major health burden worldwide, with history of infection being a significant risk factor. While the gut is a known reservoir for uropathogenic bacteria, the role of the microbiota in rUTI remains unclear. We conducted a year-long study of women with (n = 15) and without (n = 16) history of rUTI, from whom we collected urine, blood and monthly faecal samples for metagenomic and transcriptomic interrogation. During the study 24 UTIs were reported, with additional samples collected during and after infection. The gut microbiome of individuals with a history of rUTI was significantly depleted in microbial richness and butyrate-producing bacteria compared with controls, reminiscent of other inflammatory conditions. However, Escherichia coli gut and bladder populations were comparable between cohorts in both relative abundance and phylogroup. Transcriptional analysis of peripheral blood mononuclear cells revealed expression profiles indicative of differential systemic immunity between cohorts. Altogether, these results suggest that rUTI susceptibility is in part mediated through the gut–bladder axis, comprising gut dysbiosis and differential immune response to bacterial bladder colonization, manifesting in symptoms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Metagenomic sequence data are available from the Sequence Read Archive under Bioproject PRJNA400628. PBMC RNA-seq data are available from the database of Genotypes and Phenotypes (dbGaP) under project no. phs002728. Questionnaire data and output files from MetaPhlan2, Humann2 and StrainGE are available from github.com/cworby/UMB-study. Source data are provided with this paper.
Code availability
Custom R scripts used to analyse outputs are available from github.com/cworby/UMB-study.
References
Flores-Mireles, A. L., Walker, J. N., Caparon, M. & Hultgren, S. J. Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 13, 269–284 (2015).
Hooton, T. M. et al. A prospective study of risk factors for symptomatic urinary tract infection in young women. N. Engl. J. Med. 335, 468–474 (1996).
Yamamoto, S. et al. Genetic evidence supporting the fecal-perineal-urethral hypothesis in cystitis caused by Escherichia coli. J. Urol. 157, 1127–1129 (1997).
Nielsen, K. L., Dynesen, P., Larsen, P. & Frimodt-Moller, N. Faecal Escherichia coli from patients with E. coli urinary tract infection and healthy controls who have never had a urinary tract infection. J. Med. Microbiol. 63, 582–589 (2014).
Jantunen, M. E., Saxen, H., Lukinmaa, S., Ala-Houhala, M. & Siitonen, A. Genomic identity of pyelonephritogenic Escherichia coli isolated from blood, urine and faeces of children with urosepsis. J. Med. Microbiol. 50, 650–652 (2001).
Magruder, M. et al. Gut uropathogen abundance is a risk factor for development of bacteriuria and urinary tract infection. Nat. Commun. 10, 5521 (2019).
Thänert, R. et al. Comparative genomics of antibiotic-resistant uropathogens implicates three routes for recurrence of urinary tract infections. mBio 10, e01977-19 (2019).
Paalanne, N. et al. Intestinal microbiome as a risk factor for urinary tract infections in children. Eur. J. Clin. Microbiol. Infect. Dis. 37, 1881–1891 (2018).
Magruder, M. et al. Gut commensal microbiota and decreased risk for Enterobacteriaceae bacteriuria and urinary tract infection. Gut Microbes 12, 1805281 (2020).
Tariq, R. et al. Fecal microbiota transplantation for recurrent Clostridium difficile infection reduces recurrent urinary tract infection frequency. Clin. Infect. Dis. 65, 1745–1747 (2017).
Wang, T., Kraft, C. S., Woodworth, M. H., Dhere, T. & Eaton, M. E. Fecal microbiota transplant for refractory Clostridium difficile infection interrupts 25-year history of recurrent urinary tract infections. Open Forum Infect. Dis. 5, ofy016 (2018).
Mayer, E. A., Tillisch, K. & Gupta, A. Gut/brain axis and the microbiota. J. Clin. Invest. 125, 926–938 (2015).
Cryan, J. F. et al. The microbiota-gut-brain axis. Physiol. Rev. 99, 1877–2013 (2019).
Budden, K. F. et al. Emerging pathogenic links between microbiota and the gut-lung axis. Nat. Rev. Microbiol. 15, 55–63 (2017).
Dang, A. T. & Marsland, B. J. Microbes, metabolites, and the gut-lung axis. Mucosal Immunol. 12, 843–850 (2019).
Lazar, V. et al. Aspects of gut microbiota and immune system interactions in infectious diseases, immunopathology, and cancer. Front. Immunol. 9, 1830 (2018).
Scholes, D. et al. Risk factors for recurrent urinary tract infection in young women. J. Infect. Dis. 182, 1177–1182 (2000).
Clemente, J. C., Manasson, J. & Scher, J. U. The role of the gut microbiome in systemic inflammatory disease. Brit. Med. J. 360, j5145 (2018).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Parada Venegas, D. et al. Short chain fatty acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front. Immunol. 10, 277 (2019).
Liu, H. et al. Butyrate: a double-edged sword for health? Adv. Nutr. 9, 21–29 (2018).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968 (2018).
Mack, A. et al. Changes in gut microbial metagenomic pathways associated with clinical outcomes after the elimination of malabsorbed sugars in an IBS cohort. Gut Microbes 11, 620–631 (2020).
Palleja, A. et al. Recovery of gut microbiota of healthy adults following antibiotic exposure. Nat. Microbiol. 3, 1255–1265 (2018).
Zaura, E. et al. Same exposure but two radically different responses to antibiotics: resilience of the salivary microbiome versus long-term microbial shifts in feces. mBio 6, e01693-15 (2015).
Rooney, A. M. et al. Each additional day of antibiotics is associated with lower gut anaerobes in neonatal intensive care unit patients. Clin. Infect. Dis. 70, 2553–2560 (2020).
Schubert, A. M. et al. Microbiome data distinguish patients with Clostridium difficile infection and non-C. difficile-associated diarrhea from healthy controls. mBio 5, e01021-14 (2014).
Pozuelo, M. et al. Reduction of butyrate- and methane-producing microorganisms in patients with irritable bowel syndrome. Sci. Rep. 5, 12693 (2015).
Geirnaert, A. et al. Butyrate-producing bacteria supplemented in vitro to Crohn’s disease patient microbiota increased butyrate production and enhanced intestinal epithelial barrier integrity. Sci. Rep. 7, 11450 (2017).
Ni, J., Wu, G. D., Albenberg, L. & Tomov, V. T. Gut microbiota and IBD: causation or correlation? Nat. Rev. Gastroenterol. Hepatol. 14, 573–584 (2017).
Schaubeck, M. et al. Dysbiotic gut microbiota causes transmissible Crohn’s disease-like ileitis independent of failure in antimicrobial defence. Gut 65, 225–237 (2016).
Integrative Human Microbiome Project Research Network Consortium. The Integrative Human Microbiome Project: dynamic analysis of microbiome-host omics profiles during periods of human health and disease. Cell Host Microbe 16, 276–289 (2014).
Zhou, Y. & Zhi, F. Lower level of Bacteroides in the gut microbiota is associated with inflammatory bowel disease: a meta-analysis. BioMed. Res. Int. 2016, 5828959 (2016).
Duvallet, C., Gibbons, S. M., Gurry, T., Irizarry, R. A. & Alm, E. J. Meta-analysis of gut microbiome studies identifies disease-specific and shared responses. Nat. Commun. 8, 1784 (2017).
Asnicar, F. et al. Blue poo: impact of gut transit time on the gut microbiome using a novel marker. Gut 70, 1665–1674 (2021).
Takahashi, D. et al. Microbiota-derived butyrate limits the autoimmune response by promoting the differentiation of follicular regulatory T cells. EBioMedicine 58, 102913 (2020).
Rosser, E. C. et al. Microbiota-derived metabolites suppress arthritis by amplifying aryl-hydrocarbon receptor activation in regulatory B cells. Cell Metab. 31, 837–851(2020).
Li, F., Wang, M., Wang, J., Li, R. & Zhang, Y. Alterations to the gut microbiota and their correlation with inflammatory factors in chronic kidney disease. Front. Cell. Infect. Microbiol. 9, 206 (2019).
Adar, T., Shteingart, S., Ben Ya’acov, A., Bar-Gil Shitrit, A. & Goldin, E. From airway inflammation to inflammatory bowel disease: eotaxin-1, a key regulator of intestinal inflammation. Clin. Immunol. 153, 199–208 (2014).
Adar, T. et al. The importance of intestinal eotaxin-1 in inflammatory bowel disease: new insights and possible therapeutic implications. Dig. Dis. Sci. 61, 1915–1924 (2016).
Cheung, W., Bluth, M., Khan, S., Johns, C. & Bluth, M. Peripheral blood mononuclear cell gene array profiles in female patients with involuntary bladder contractions. Adv. Genomics Genet. 1, 3–7 (2011).
de Santiago, P. R. et al. Immune-related IncRNA LINC00944 responds to variations in ADAR1 levels and it is associated with breast cancer prognosis. Life Sci. 268, 118956 (2021).
Gur, C. et al. Natural killer cell-mediated host defense against uropathogenic E. coli is counteracted by bacterial hemolysinA-dependent killing of NK cells. Cell Host Microbe 14, 664–674 (2013).
Rivera-Chavez, F. et al. Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host Microbe 19, 443–454 (2016).
Antharam, V. C. et al. Intestinal dysbiosis and depletion of butyrogenic bacteria in Clostridium difficile infection and nosocomial diarrhea. J. Clin. Microbiol. 51, 2884–2892 (2013).
van Dijk, L. et al. StrainGE: a toolkit to track and characterize low-abundance strains in complex microbial communities. Genome Biol. 23, 74 (2022).
Clermont, O., Bonacorsi, S. & Bingen, E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl. Environ. Microbiol. 66, 4555–4558 (2000).
Schreiber, H. L. et al. Bacterial virulence phenotypes of Escherichia coli and host susceptibility determine risk for urinary tract infections. Sci. Transl. Med. 9, eaaf1283 (2017).
Garretto, A. et al. Genomic survey of E. coli from the bladders of women with and without lower urinary tract symptoms. Front. Microbiol. 11, 2094 (2020).
Zhang, S. et al. Short chain fatty acids modulate the growth and virulence of pathosymbiont Escherichia coli and host response. Antibiotics (Basel) 9, 462 (2020).
Stapleton, A. E. The vaginal microbiota and urinary tract infection. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.uti-0025-2016 (2016).
Forde, B. M. et al. Population dynamics of an Escherichia coli ST131 lineage during recurrent urinary tract infection. Nat. Commun. 10, 3643 (2019).
Cusumano, C. K. et al. Treatment and prevention of urinary tract infection with orally active FimH inhibitors. Sci. Transl. Med. 3, 109ra115 (2011).
Dethlefsen, L. & Relman, D. A. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc. Natl Acad. Sci. USA 108, 4554–4561 (2011).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Acknowledgements
This project has been funded in part with Federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services under grant no. U19AI110818 to the Broad Institute, from the National Institutes of Health Mucosal Immunology Studies Team consortium under grant no. U01AI095542 to Washington University and the National Institute of Diabetes and Digestive and Kidney Disease, National Institutes of Health, Department of Health and Human Services, under Grant Number R01DK121822 to the Broad Institute and Washington University. B.S.O. was supported by grants from the National Institutes of Health, USA (nos. T32GM007067 and T32GM139774). A.L.K. was supported by grants from the National Institutes of Health, Department of Health, USA (no. R01AI165915) and the Doris Duke Charitable Foundation. This work was also supported by funds from the Center for Women’s Infectious Disease Research at Washington University School of Medicine. We thank members of the Broad’s Bacterial Genomics group and H. Vlamakis for helpful conversations. We thank B. Haas for assistance with PBMC RNA-seq analysis, as well as the Multi-Omics Core and Genomics Platform at the Broad Institute for sample processing and data generation.
Author information
Authors and Affiliations
Contributions
Study design was undertaken by H.L.S., K.W.D., S.J.H. and A.M.E. Study coordination was carried out by H.L.S., K.B., S.B.C. and A.K. Experiments were performed by H.L.S., J.S.P., C.L.P.O., V.L.M. and A.E.P. Data analysis was undertaken by C.J.W., H.L.S., T.J.S., L.R.v.D., R.A.B., B.S.O., B.J.H., C.A.D. and W.-C.C. Consultation and supervision of analyses were the responsibility of B.J.W., A.L.M., T.J.H., T.M.H., A.L.K., H.H.L., K.W.D., S.J.H. and A.M.E. C.J.W., A.L.M., K.W.D., S.J.H. and A.M.E. prepared the original draft. Review and approval of the final manuscript was provided by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks John Lee, Alice McHardy and Mark Schembri for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sex precedes all clinical UTI events.
Survey reports of intercourse frequency in the previous two weeks. Responses are partitioned by (i) control women, (ii) rUTI women at time of UTI, and (iii) rUTI women at non-UTI time points.
Extended Data Fig. 2 SCFA producing bacteria are depleted in the rUTI gut.
Cumulative relative abundances of (a) butyrate and (b) propionate producing bacterial species in rUTI and control samples. Box plots display the median (center line), 25th and 75th percentiles (box), as well as the 5th and 95th percentiles (whiskers). Within-host average relative abundances of individual species for (c) butyrate and (d) propionate producers are also shown. Horizontal lines denote the mean relative abundance in rUTI (red) and control (blue) women.
Extended Data Fig. 3 Bray Curtis dissimilarity across stool samples.
(a) For each patient, the distribution of Bray-Curtis dissimilarities between all stool samples, ordered by increasing mean patient values within each cohort. (b) Bray-Curtis distributions between samples taken at the time of UTI vs. healthy time points (red), compared to all pairwise healthy sample comparisons. Box plots show the median (center line), 25th and 75th percentiles (box), as well as the 5th and 95th percentiles (whiskers).
Extended Data Fig. 4 rUTI dysbiosis is not driven by antibiotic use during the study.
We grouped rUTI women according to their antibiotic exposures at any point during the UMB study; (i) ciprofloxacin (n = 6) (ii) non-ciprofloxacin antibiotics (n = 6); (iii) no antibiotics (n = 3); (iv) any antibiotics (n = 12). Groups were compared against each other and against the control cohort (n = 16) for (a) overall microbial richness and (b) relative abundance of butyrate producers. Crosses represent mean values for individuals, boxplots denote the IQR and 95% central quantiles for each group. Wilcoxon rank sum tests (two-sided) were applied to group pairs to derive p-values. (c) Temporal trends of microbial richness (black) and relative abundance of butyrate producers (red) in all rUTI participants using antibiotics during the study. For each individual, linear models were fit to observations (points) over time; fitted trends are shown, with coefficients & p values reported at the top of each panel. Dashed vertical lines denote antibiotic usage. Participant mean values are represented by horizontal lines.
Extended Data Fig. 5 Most species depleted in the rUTI gut are also depleted in the IBD gut.
We compared discriminatory taxa in rUTI women to those in IBD patients using data from adult participants in the HMP2 study33. For each study, we fitted mixed effects models to standardized Metaphlan2 relative abundances as a function of categorical disease group (rUTI or IBD respectively, vs. each study’s control cohort), including covariates for race and antibiotic use. The disease group coefficients are plotted against each other for each species, with circle pairs representing the average relative abundance in each study. Species with uncorrected p values <0.05 in either study are labelled. Species not present in at least 10% of samples in either study are excluded. IBD comprises patients with either CD or UC.
Extended Data Fig. 6 Immunological differences between cohorts.
(a) PCA plot of gene expression across cohorts, based on PBMC RNA Seq data. Samples are partitioned into healthy controls (n = 13), rUTI patient baseline (enrollment; n = 12) and rUTI patient at time of UTI (n = 17). (b) Plasma eotaxin-1 levels in control women, and rUTI women at healthy enrollment and time of UTI. (c) Relative abundance of NK cells in control and rUTI women based on CIBERSORT output. Box plots display the median (center line), 25th and 75th percentiles (box), as well as data points within 1.5 IQR of the upper & lower quartiles (whiskers), and outliers beyond this range (dots).
Extended Data Fig. 7 Limited relationship between non SCFA-producing taxa with butyrate producers.
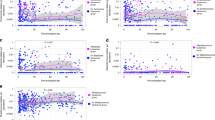
For all non SCFA-producing genera detected across all samples, the correlation coefficient between its relative abundance and the relative abundance of butyrate producers was calculated and plotted against its mean relative abundance across (a) control (n = 170) and (b) rUTI (n = 197) samples. Genera with an absolute correlation coefficient greater than 0.25 are labeled, along with Escherichia, represented by the red point.
Extended Data Fig. 8 E. coli relative abundance around the time of UTI and phylogroup distributions.
For all stool samples taken within 3 days of a UTI event, the log fold change is given relative to (a) the median E. coli relative abundance in the corresponding patient, excluding samples taken at the time of UTI, and (b) the relative abundance of E. coli in the preceding stool sample. ‘X’ denotes samples for which there was no prior sample available. (c) Number of detected E. coli strains by sample type. (d) Number of detected StrainGST reference strains vs. relative abundance of E. coli.
Extended Data Fig. 9 Strain dynamics in control women.
Strain dynamics within all control participants; analogous to Fig. 3. (a) Phylogenetic tree comprising strains called by StrainGE across all stool and urine samples, colored by phylogroup. Bars show number of unique participants with at least one strain observation; bars are bolded if the strain was identified in at least one urine sample. Each strain identified in control women is uniquely identifiable by the phylogroup (colour) and ID (numeral) indicated right. (b) Each panel represents longitudinal strain dynamics within one patient. Numerals refer to strain identifiers in (a). All fecal strains are connected to their most recent previous observation in fecal samples. Diamonds denote clinical rectal swabs. Strains identified in urine outgrowth depicted if available; otherwise raw urine strains are shown. Fecal or urine samples with no detected E. coli strains represented by open grey symbols. Vertical dashed lines represent self-reported antibiotic use.
Supplementary information
Supplementary Tables
Supplementary Data Tables 1–9.
Source data
Source Data Fig. 1
Statistical Source data.
Source Data Fig. 2
Statistical Source data.
Source Data Fig. 3
Statistical Source data.
Source Data Fig. 4
Statistical Source data.
Source Data Extended Data Fig. 1
Statistical Source data.
Source Data Extended Data Fig. 2
Statistical Source data.
Source Data Extended Data Fig. 3
Statistical Source data.
Source Data Extended Data Fig. 4
Statistical Source data.
Source Data Extended Data Fig. 5
Statistical Source data.
Source Data Extended Data Fig. 6
Statistical Source data.
Source Data Extended Data Fig. 7
Statistical Source data.
Source Data Extended Data Fig. 8
Statistical Source data.
Source Data Extended Data Fig. 9
Statistical Source data.
Rights and permissions
About this article
Cite this article
Worby, C.J., Schreiber, H.L., Straub, T.J. et al. Longitudinal multi-omics analyses link gut microbiome dysbiosis with recurrent urinary tract infections in women. Nat Microbiol 7, 630–639 (2022). https://doi.org/10.1038/s41564-022-01107-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01107-x
This article is cited by
-
Fabrication of levofloxacin-loaded porcine acellular dermal matrix hydrogel and functional assessment in urinary tract infection
Journal of Nanobiotechnology (2024)
-
Microbiota–gut–brain axis and its therapeutic applications in neurodegenerative diseases
Signal Transduction and Targeted Therapy (2024)
-
Evaluating occult causes of disease: the tricompartmental PNEI approach and the importance of the microbiome
Techniques in Coloproctology (2024)
-
Dysbiosis of a microbiota–immune metasystem in critical illness is associated with nosocomial infections
Nature Medicine (2023)
-
Human microbiome myths and misconceptions
Nature Microbiology (2023)